Tag: Genome-wide Association Studies
-
Tutorial on Polygenic Risk Score
This note is based on Choi, S. W., Mak, T. S.-H., & O’Reilly, P. F. (2020). Tutorial: A guide to performing polygenic risk score analyses. Nature Protocols, 15(9), Article 9.
- LD Score Regression
- Joint Local False Discovery Rate in GWAS
-
Quantitative Genetics
This post is based on the Pao-Lu Hsu Award Lecture given by Prof. Hongyu Zhao at the 11th ICSA International Conference on Dec. 21th, 2019.
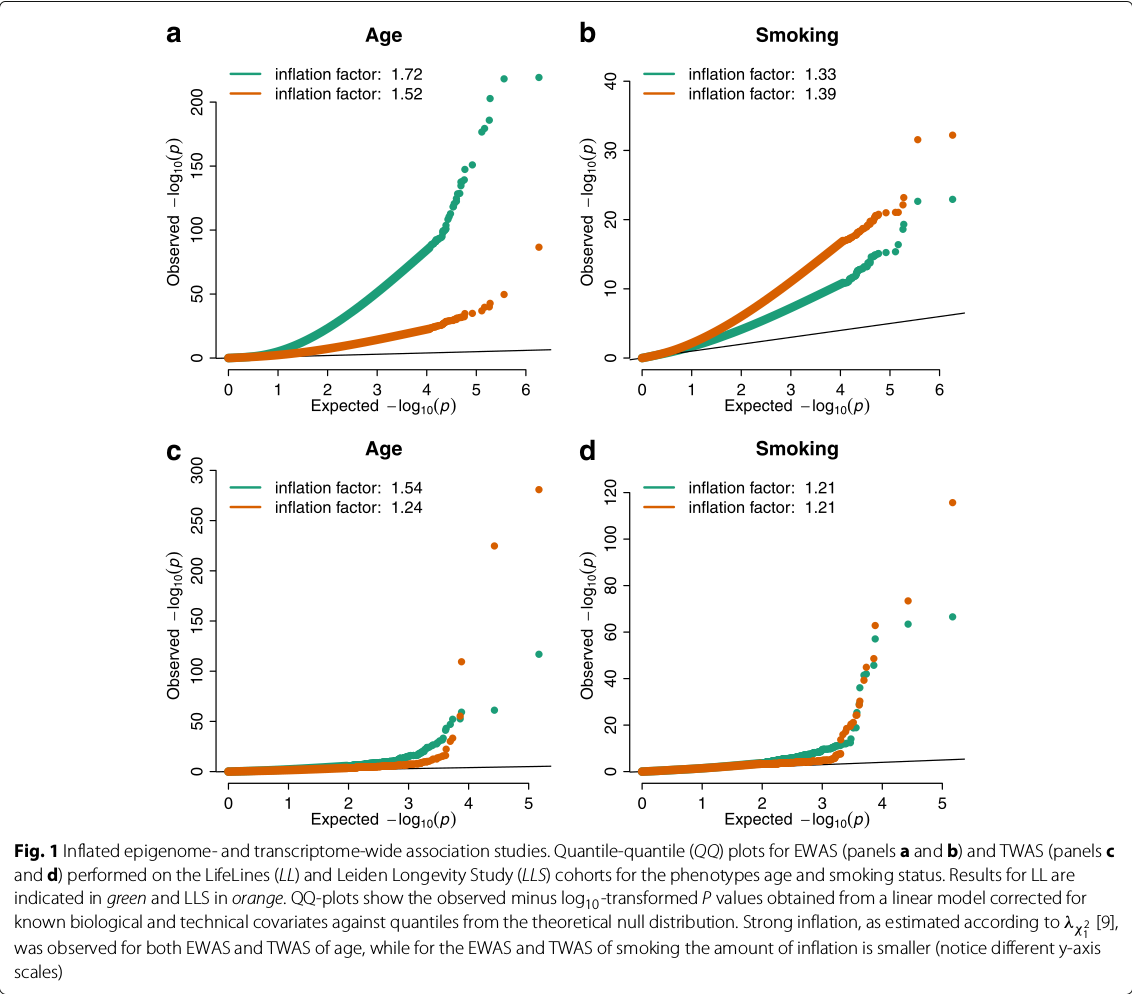
- Controlling bias and inflation in EWAS/TWAS
-
Genetic Relatedness in High-Dimensional Linear Models
This post is based on Guo, Z., Wang, W., Cai, T. T., & Li, H. (2019). Optimal Estimation of Genetic Relatedness in High-Dimensional Linear Models. Journal of the American Statistical Association, 114(525), 358–369.
Archive
3D Registration2
Adaptive Algorithm2
Additive Model1
Admissible2
Annealing2
Approximate Bayesian Computation1
Approximation1
Astronomy1
Asymptotic2
B-spline3
Baum-Welch Algorithm1
Bayes Factor1
Bayes Risk1
Bayesian Analysis1
Bayesian Inference7
Bayesian Model1
Bayesian Shrinkage1
Bernstein1
Berry-Esseen Bound1
Biclustering1
Big Data1
Binomial Thinning1
Biobank1
Bioinformatics5
Biomarkers1
Biostatistics1
Bipartite Graph2
Bipartitle Graph1
Biplot1
Bootstrap5
Bridge Sampling1
C-index1
CNN1
Canonical1
Causal Inference3
Cell Cycle1
Cell Tracking1
Cell Type1
Central Limit Theory1
Chain Structure1
Change Points2
Chinese Medical Conversations1
Chinese Restaurant Process1
Chinese Word Segmentation2
Classification2
Classification Tree1
Clustering4
Common Principal Component Analysis1
Composite Likelihood1
Compressed Sensing1
Concentration Inequality2
Conditional Distribution1
Cone Constraint1
Confidence Interval1
Confidence Intervals2
Conformal Inference2
Conjugate Gradient2
Consistency1
Continuous Time Markov Chain3
Contrastive Learning2
Copula2
Copy Number Variation1
Cosine Embeddings1
Coupled Clustering1
Covariance Estimation1
Covariance Function1
Covariance Matrix Estimation1
Covariance Test1
Cox Model5
Cross-Validation7
DNA Methylation2
Dantzig1
Data Alignment2
Data Augmentation1
Data Perturbation1
Debiased1
Debiased Lasso2
Deep Learning6
Deep Neural Networks1
Degrees of Freedom2
Delta Method2
Differential Equations1
Differential Expression7
Differential privacy1
Dimension Reduction2
Directed Graph1
Discrete Time Markov Chain1
Discretization1
Disease Subtyping1
Dispersion1
Distributed Learning1
Divide-and-Conquer2
Dropout1
Dynamic Programming1
EHR1
ELBO2
EM Algorithm1
Economy2
Edgeworth1
Effective Sample Size1
Eigenvalues1
Electronic Health Records2
Electronic Medical Records1
Embedding2
Empirical Bayes3
Empirical Bayes Information1
Empirical Process1
Environmental Stochasticity1
Epidemic1
Epigenome-wide Association Studies1
Epistasis1
Equicorrelation1
Equivariance2
Estimability1
Estimating Equations1
Evolution2
Exchangeability1
FDR2
Factor Models1
False Discovery Rate7
Feature Selection1
Forward Stepwise Regression1
Forward-Backward Algorithm1
Fourier1
Functional Data Analysis9
GWAS5
Gaussian Processes1
Gaussian Quadrature1
Gene Expression1
Gene Regulatory Network1
Generalization1
Generalized Additive Model1
Generalized Linear Model3
Generalized Linear Regression1
Generative Pre-trained Transformer1
Generator1
Genetic Fine Mapping2
Genome-wide Association Studies6
Gibbs4
Gradient Descent2
Graph7
Graph Neural Network1
Graph Partition1
Group Invariance Test1
Group Lasso1
Group Testing1
Hammersley-Clifford Theorem1
Helicobacter pylori2
Heritability1
Hidden Markov Model3
Hierarchical Clustering4
Hierarchical Model3
Hierarchical Multi-label Classification2
High-Dimensional12
Horseshoe1
Hypothesis Testing7
IPF3
Identifiability1
Imbalanced Classification1
Importance Sampling6
In-context Learning1
Infinite Relational Model1
Information Matrix1
Instance Segmentation2
Integration Study1
Interpolation3
Intersection-union Test2
Invariance2
Isotonic Regression3
Item Response Theory1
Iterative Closest Point3
Jackknife2
James-Stein3
Jeffreys1
Joint Model1
K-means2
KEGG1
KKT Conditions1
KL-divergence1
Kalman Filter2
Kernel Association Test2
Kernel Ridge Regression2
Key-value memory networks1
Knockoff3
Knockoffs1
Knowledge Graph1
Krylov Space1
LR Test1
Lady Tasting Tea1
Lagrange Multiplier1
Lasso12
Least Angle Regression1
Least Squares Estimators1
Levenberg-Marquardt1
Likelihood1
Likelihood-free Inference1
Linear Assignment1
Linear Assignment Problem1
Linear Discriminant Analysis4
Linear Mixed-Effects1
Linear Model2
Linear Programming4
Linear Regression8
Linear Smoothers1
Linkage Disequilibrium1
Location-Scale Family2
Logistic Regression2
Longitudinal2
M-estimator1
MAP1
MLE4
Machine Learning1
Magnetic Field1
Manifold Learning1
Marginal Distribution1
Marginal Likelihood1
Markov Random Field2
Mask Embeddings1
Matrix Completion3
Mediation Analyses1
Medical Concept Recognition1
Mendelian Randomization1
Meta-analysis2
Metabolic Network4
Metropolis Hastings1
Microarray2
Microbiome2
Minimax7
Minimum-Cost Flow2
Minorization-maximization1
Mixed-Models1
Mixed-effects Model1
Mixture Model6
Model Selection6
Monotone Function8
Monotone Regression1
Monte Carlo1
Monte Carlo Integration1
Multi-omics4
Multidimensional Scaling1
Multiple Object Tracking7
Multivariate1
Multivariate Adaptive Regression Splines1
Mutual Incoherence1
Mutual Information1
NBα-distribution1
Named Entity Recognition1
Nature Language Processing5
Neural Nets1
Neural Network1
Nominal Size1
Non-central Chi-square1
Nonconvex1
Nonnegative polynomials1
Norm1
Normal1
Normalization2
Nuclear Norm2
Object Tracking4
Observational Study1
Optimal Control1
Optimization4
Overfitting1
Paradox2
Parametric Curves1
Partial Least Squares2
Particle Filter5
Path Sampling1
Penalized Splines2
Permutation Test3
Phylogeny3
Poisson1
Poisson Regression1
Polygenic Risk Score2
Polygenicity1
Precision Medicine1
Presistency1
Principal Component Analysis9
Principal Curves3
Prior1
Probability1
Probability Measure2
Propensity Score1
Protein Folding1
Proximal Operator1
Pseudolikelihood3
Pseudotime8
Quantile Curves1
Quantile Regression3
Quasi-likelihood2
Quaternion1
RJMCMC1
RKHS1
RNN1
ROC1
Rademacher Complexity1
Radiology1
Random Effects2
Random Finite Sets1
Random Forests3
Random Matrix Theory2
Randomization Test1
Rank1
Rao-Blackwell1
Rare Variant Association1
Reaction Network1
Recommendation System1
Record Linkage1
Recurrent Hourglass Networks1
Registration1
Regression Tree4
Rejection Method1
Resampling1
Restricted Eigenvalues1
Restricted Isometry1
Restricted Maximum Likelihood3
Ridge5
Risk Function1
SGD1
SMC-PHD2
SVM1
Scale Parameter3
Score Test1
Second-order Programming1
Selection Bias1
Selective Inference5
Self-avoid Walk1
Self-normalized1
Self-organized Map1
Semiparametric1
Sequence Alignment1
Sequencing1
Sequential Importance Sampling2
Sequential Monte Carlo9
Shallow Learners1
Shape Restrictions1
Shape-constrained4
Shuffled Data1
Single Index Model2
Single-cell23
Singular Value Decomposition1
Sliced Inverse Regression4
Smoothing Splines3
Soft Threshold1
Sort1
Spacing Test1
Sparse Group Lasso1
Sparsistency2
Spatial Proteomics1
Species Persistence1
Spike-and-slab Prior1
Splines1
Stan1
Standardize1
Star Formation2
Statistical Decisions1
Statistical Inference1
Statistical Learning2
Stein's Method1
Stein's Operator1
Stochastic Differential Equation1
Strong Convexity2
Studentize1
Sub-Gaussian1
Subgradient1
Sufficient Dimension Reduction1
Summary Statistics Selection1
Support Recovery1
Survey1
Survival Analysis7
Systems Biology1
Tensor1
Text Mining2
The Cancer Genome Atlas2
Time Series4
Time-varying5
Toeplitz1
Transcriptome-wide Association Studies1
Transformer2
Transformers1
Tree Variability2
Tuberculosis1
Tumor Heterogeneity1
Tweedie1
Two-phase Regression1
U-Statistics2
Uniform Convergence1
Union-intersection Test1
Variable Selection9
Variance Component Analysis1
Variance Components Model1
Variance-Component Score Test1
Variational Inference5
Varying-Coefficient1
Viterbi Algorithm1
Wald Test1
Wasserstein Space1
Woodbury Formula2
XGBoost1
eQTL2
p-value2
p-values2