Bootstrap Sampling Distribution
Posted on (Update: )
This note is based on Lehmann, E. L., & Romano, J. P. (2005). Testing statistical hypotheses (3rd ed). Springer.
Let $X^n = (X_1,\ldots, X_n)$ be a sample of $n$ i.i.d. random variables taking values in a sample space $S$ and having unknown probability distribution $P\in \cP$. The interest is to infer some parameter $\theta(P)$. Denote $\Theta = \{\theta(P):P\in\cP\}$ be the range of $\theta$.
Consider a term called root, $R_n(X^n, \theta(P))$, which is just some real-valued functional depending on both $X^n$ and $\theta(P)$. The idea is that a CI for $\theta(P)$ could be constructed if the distribution of the root were known.
If the problem is nonparametric, a natural construction for an estimator $\hat\theta_n$ of $\theta(P)$ is the plug-in estimator $\hat\theta_n=\theta(\hat P_n)$, where $\hat P_n$ is the empirical distribution of the data,
\[\hat P_n(E) = \frac 1n\sum_{i=1}^nI\{X_i\in E\}\,.\]For parametric problem, i.e., $\cP = \{P_\psi:\psi\in \Psi\}$, then $\theta(P)$ can be described as a functional $t(\psi)$, and hence $\hat\theta_n$ is often taken to be $t(\hat\psi_n)$, where $\hat\psi_n$ is some desirable estimator of $\psi$.
Let $J_n(P)$ be the distribution of $R_n(X^n, \theta(P))$ under $P$, and let $J_n(\cdot, P)$ be the corresponding CDF defined by
\[J_n(x, P) = P\{R_n(X^n, \theta(P))\le x\}\,.\]The bootstrap procedure is a general, direct approach to approximate the sampling distribution $J_n(P)$ by $J_n(\hat P_n)$, where $\hat P_n$ is an estimate of $P$ in $\cP$. In this light, the bootstrap estimate $J_n(\hat P_n)$ is a simple plug-in estimate of $J_n(P)$.
- in parametric estimation, $\hat P_n$ maybe $P_{\hat \psi_n}$
- in nonparametric estimation, $\hat P_n$ is typically the empirical distribution
In order to get around the non-continuity and non-strictly-increasing problem, define
\[J_n^{-1} = \inf\{x:J_n(x,P)\ge 1-\alpha\}\,,\]then if $J_n(\cdot, P)$ has a unique quantile $J_n^{-1}(1-\alpha, P)$,
\[P(R_n(X^n, \theta(P))\le J_n^{-1}(1-\alpha, P)) = 1-\alpha\,.\]The resulting bootstrap CI for $\theta(P)$ of nominal level $1-\alpha$ could be
\[\{\theta\in \Theta:R_n(X^n,\theta) \le J_n^{-1}(1-\alpha, \hat P_n)\}\]or
\[\{\theta\in \Theta:J_n^{-1}(\alpha/2, \hat P_n)\le R_n(X^n,\theta) \le J_n^{-1}(1-\alpha/2, \hat P_n)\}\]Outside certain exceptional cases (TODO), the bootstrap approximation $J_n(x,\hat P_n)$ cannot be calculated exactly.
Even in the relatively simple case when $\theta(P)$ is the mean of $P$, the root is $n^{1/2}[\bar X_n - \theta(P)]$, and $\hat P_n$ is the empirical distribution, the exact computation of the bootstrap distribution involves an n-fold convolution.
Typically, we resort to a Monte Carlo approximation to $J_n(P)$, conditional on the data $X^n$, for $j=1,\ldots,B$, let $X_j^{n*}$ be a sample of $n$ i.i.d. observation from $\hat P_n$, which is also referred to as the $j$-th bootstrap sample of size $n$. When $\hat P_n$ is the empirical distribution, this amounts to resampling the original observations with replacement.
Consider the consistency of the bootstrap estimator $J_n(\hat P_n)$ of the true sampling distribution $J_n(P)$ of $R_n(X^n,\theta(P))$. For the bootstrap to be consistent, $J_n(P)$ must be smooth in $P$ since we are replacing $P$ by $\hat P_n$.

Parametric Bootstrap
Suppose $X^n=(X_1,\ldots, X_n)$ is a sample from a q.m.d. model $\{P_\theta,\theta\in \Omega\}$.
The family $\{P_\theta,\theta\in \Omega\}$ is quadratic mean differentiable (q.m.d.) at $\theta_0$ if there exists a vector of real-valued functions $\eta(\cdot,\theta_0) = (\eta_1(\cdot,\theta_0),\ldots,\eta_k(\cdot,\theta_0))^T$ such that
\[\int_\cX\left[\sqrt{p_{\theta_0+h}(x)} -\sqrt{p_{\theta_0}(x)}-\langle \eta(x,\theta_0),h\rangle\right]^2d\mu(x) = o(\vert h\vert^2)\]as $\vert h\vert\rightarrow 0$.
Suppose $\hat \theta_n$ is an efficient likelihood estimator in the sense that
\[n^{1/2}(\hat\theta_n-\theta_0) = I^{-1}(\theta_0)Z_n + o_{P_{\theta_0}^n}(1)\,,\]where $Z_n$ is the normalized score vector
\[Z_n = Z_n(\theta_0) = 2n^{-1/2}\sum_{i=1}^n[\eta(X_i,\theta_0)/p_{\theta_0}^{1/2}(X_i)]\,.\]Suppose $g(\theta)$ is a differentiable map from $\Omega$ to $\IR$ with nonzero graident vector $\dot g(\theta)$. Consider the root $R_n(X^n,\theta) = n^{1/2}[g(\hat\theta_n)-g(\theta)]$, with distribution function $J_n(x,\theta)$. It can show that
\[J_n(x,\theta)\rightarrow N(0, \dot g(\theta)I^{-1}(\theta)\dot g(\theta)^T) \triangleq N(0, \sigma_\theta^2)\,.\]It may difficult to calculate the limiting variance $\sigma_\theta^2$, then one can use the approximation
\[\hat\sigma_n^2 = \dot g(\hat\theta_n) I^{-1}(\hat\theta_n)\dot g(\hat \theta_n)^T\,,\]which can be poor. Then we can consider bootstrap method, use the approximation $J_n(x,\hat\theta_n)$, which is differ from the above $\widehat{\lim J_n(x,\theta)}$.
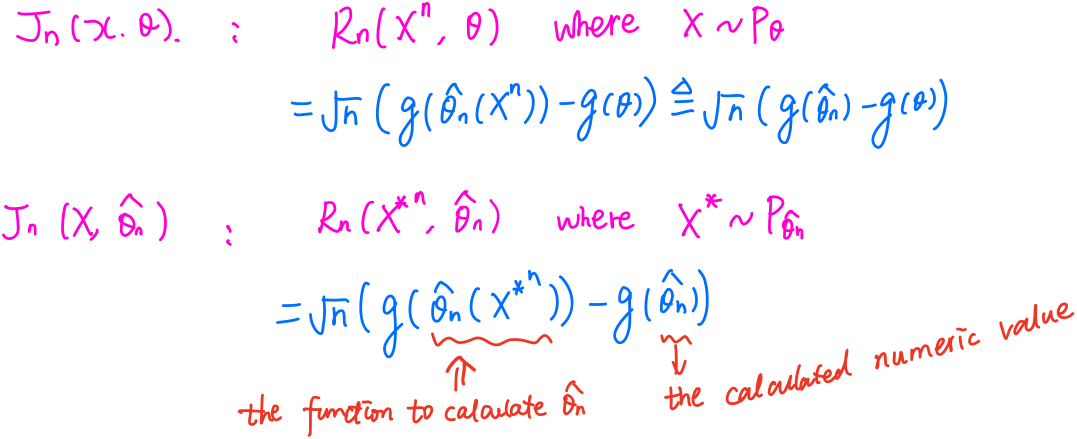
Let $X_1,\ldots, X_n$ be i.i.d. random variables with pdf $\frac 1\sigma f(\frac{x-\mu}{\sigma})$, where $f$ is a known Lebesgue pdf and $\mu,\sigma$ are unknown. Let $X_1^*,\ldots, X_n^*$ be i.i.d. bootstrap data from the pdf $\frac 1s f(\frac{x-\bar x}{s})$, where $\bar x$ and $s^2$ are the observed sample mean and sample variance, respectively.
Higher Order Asymptotic Comparisons
One of the main reasons the bootstrap approach is so valuable is that it can be applied to approximate the sampling distribution of an estimator in situations where the finite or large sample distribution theory is intractable, or depends on unknown parameters
Even in relatively simple situations, there are advantages to using a bootstrap approach. For example, consider the problem of constructing a confidence interval for a mean. Under the assumption of a finite variance, the standard normal theory interval and the bootstrap-t are each pointwise consistent in level. In order to compare them, we must consider higher order asymptotics properties.
More generally, suppose $I_n$ is a nominal $1 - \alpha$ level confidence interval for a parameter $\theta(P)$. Its converge error under $P$ is
\[P(\theta(P) \in I_n) - (1 - \alpha)\]and we would like to examine the rate at which this tends to zero.
In typical problems, this coverage error is a power of $n^{-1/2}$. It will be necessary to distinguish one-sided and two-sided confidence intervals because their orders of coverage error may differ.
In this section, we focus on confidence intervals for the mean in a nonparameteric setting. Compare some asymptotic methods based on the normal approximation and the bootstrap.
Let $X^n = (X_1,\ldots, X_n)$ be i.i.d. with c.d.f. $F$, mean $\theta(F)$, and variance $\sigma^2(F)$.
Also, let $\hat F_n$ denote the empirical c.d.f., and let $\hat\sigma_n = \sigma(\hat F_n)$
Consider the roots
\[R_n(X^n, F) = n^{1/2}(\bar X_n - \theta(F))\]and
\[R_n^s(X^n, F) = n^{1/2}(\bar X_n - \theta(F)) / \hat\sigma_n\]Their distribution functions under $F$ are denoted $J_n(\cdot, F)$ and $K_n(\cdot, F)$, respectively.
Assume $E_F(X_i^4) < \infty$. Let $\psi_F$ denote the characteristic function of $F$, and assume \(\limsup_{\vert s\vert \rightarrow\infty} \vert\psi_F(s)\vert < 1\,.\) Then,
\[J_n(t, F) = \Phi(t/\sigma(F)) - \frac 16\gamma(F)\phi(t/\sigma(F)) \left(\frac{t^2}{\sigma^2(F)}-1\right)n^{-1/2} + O(n^{-1})\,,\tag{15.45}\]where
\[\gamma(F) = E_F[X_1 - \theta(F)]^3/\sigma^3(F)\]is the skewness of $F$. Moreover, the expansion holds uniformly in $t$ in the sense that
\(J_n(t, F) = [\Phi(t/\sigma(F)) - \frac 16\gamma(F)\psi(t/\sigma(F))(\frac{t^2}{\sigma^2(F)}-1)n^{-1/2}] + R_n(t, F)\,,\) where $\vert R_n(t, F)\vert \le C/n$ for all $t$ and some $C = C_F$ which depends on $F$
Assume $E_F(X_i^4) < \infty$ and that $F$ is absolutely continuous. Then, uniformly in $t$,
\[K_n(t, F) = \Phi(t) + \frac{1}{6} \gamma(F) \phi(t)(2t^2+1) n^{-1/2} + O(n^{-1})\tag{15.46}\]
The dominant error in using a standard normal approximation to the distribution of the studentized statistic is due to skewness of the underlying distribution.
However, note that the expansions are asymptotic results, and for finite $n$, including the correction term (the term of order $n^{-1/2}$) may worsen the approximation.
The above expansions are known as Cornish-Fisher Expansions.
$K_n^{-1}(1-\alpha, F)$ is a value of $t$ satisfying $K_n(t, F) = 1-\alpha$. Of course, $K_n^{-1}(1-\alpha, F)\rightarrow z_{1-\alpha}$.
We would like to determine $c = c(\alpha, F)$ such that
\[K_n^{-1}(1-\alpha, F) = z_{1-\alpha} + cn^{-1/2} + O(n^{-1})\]Set $1-\alpha$ equal to the right hand side of (15.46) with $t = z_{1-\alpha} + cn^{-1/2}$, which yields
\[\Phi(z_{1-\alpha} + cn^{-1/2}) + \frac 16\gamma(F) \phi(z_{1-\alpha} + cn^{-1/2})(2z_{1-\alpha}^2+1)n^{-1/2} + O(n^{-1}) = 1-\alpha\,.\]By expanding $\Phi$ and $\phi$ about $z_{1-\alpha}$, we find that
\[c = -\frac 16\gamma(F)(2z_{1-\alpha}^2 + 1)\,.\]Thus,
\[K_n^{-1}(1-\alpha, F) = z_{1-\alpha} - \frac 16 \gamma(F) (2z_{1-\alpha}^2 + 1) n^{-1/2} + O(n^{-1})\tag{15.47}\]Normal Theory Intervals
the most basic approximate upper one-sided confidence interval for the mean $\theta(F)$ is given by
\[\bar X_n + n^{-1/2}\hat\sigma_nz_{1-\alpha}\]where $\hat\sigma_n^2 = \sigma^2(\hat F_n)$ is the (biased) sample variance. Its one-sided coverage error is given by
\[P_F\{\theta(F) \le \bar X_n + n^{-1/2}\hat\sigma_nz_{1-\alpha} \} - (1-\alpha) = \alpha - P_F(n^{1/2}(\bar X_n - \theta(F))/\hat\sigma_n < z_{\alpha})\]By (15.46), the one-sided coverage error of this normal theory interval is
\[-\frac 16\gamma(F)\phi(z_\alpha)(2z_{\alpha}^2 + 1) n^{-1/2} + O(n^{-1}) = O(n^{-1/2})\]Analogously, the coverage error of the two-sided confidence interval of nominal level $1-2\alpha$,
\[\bar X_n \pm n^{-1/2} \hat\sigma_nz_{1-\alpha}\,,\]satisfies
\[P_F(-z_{1-\alpha} \le n^{1/2}(\bar X_n -\theta(F)) / \hat\sigma_n \le z_{1-\alpha}) - (1-2\alpha) = P(n^{1/2}(\bar X_n-\theta(F))/\hat\sigma_n \le z_{1-\alpha}) - P(n^{1/2}(\bar X_n -\theta(F))\hat\sigma_n < -z_{1-\alpha}) - (1-2\alpha)\]which by (15.46) is equal to
\[[\Phi(z_{1-\alpha}) + \frac 16\gamma(F)\phi(z_{1-\alpha})(2z_{1-\alpha}^2+1)n^{-1/2} + O(n^{-1})] - [\Phi(-z_{1-\alpha}) + \frac 16\gamma(F)\phi(-z_{1-\alpha})(2z_{1-\alpha}^2+1)n^{-1/2} + O(n^{-1})] - (1-2\alpha) = O(n^{-1})\]the main reason that a normal approximation is used for the distribution of $n^{1/2}(\bar X_n - \theta(F))/\hat\sigma_n$ and no correction is made for skewness of the underlying distribution.
For example, if $\gamma(F) > 0$, the one-sided upper confidence bound (15.49) undercovers slightly while the one-sided lower confidence bound overcovers.
The combination of overcoverage and undercoverage yields a net result of a reduction in the order of coverage error of two-sided intervals.
Basic Bootstrap Intervals
Consider bootstrap confidence intervals for $\theta(F)$ based on the root
\[R_n(X^n, \theta(F)) = n^{1/2} (\bar X_n - \theta(F))\]It is plausible that the bootstrap approximation $J_n(t, \hat F_n)$ to $J_n(t, F)$ satisifies an expansion like (15.45) with $F$ replaced by $\hat F_n$. In fact, it is the case that
\[J_n(t, \hat F_n) = \Phi(t/\hat\sigma_n) - \frac 16\gamma(\hat F_n)\phi(t/\hat\sigma_n) (\frac{t^2}{\hat\sigma_n^2}-1)n^{-1/2} + O_P(n^{-1})\tag{15.54}\]Both side of (15.54) are random and the remainder term is now of order $n^{-1}$ in probability.
Similarly, the bootstrap quantile function $J_n^{-1}(1-\alpha, \hat F_n)$ has an analogous expansion and is given by
\[J_n^{-1}(1-\alpha, \hat F_n) = \hat\sigma_n[z_{1-\alpha} + \frac{1}{6}\gamma(\hat F_n)(z_{1-\alpha}^2-1)n^{-1/2}] + O_P(n^{-1}) \tag{15.55}\]From (15.45) and (15.54), it follows that
\[J_n(t, \hat F_n) - J_n(t, F) = O_P(n^{-1/2})\]because
\[\hat\sigma_n - \sigma(F) = O_P(n^{-1/2})\]Consider the one-sided coverage error of the nominal level $1-\alpha$ upper confidence bound $\bar X_n - n^{-1/2}J_n^{-1}(\alpha, \hat F_n)$, given by
\[P_F(\theta(F) \le \bar X_n - n^{-1/2}J_n^{-1}(\alpha, \hat F_n)) - (1-\alpha) = \alpha - P_F(n^{1/2}(\bar X_n - \theta(F)) < J_n^{-1}(\alpha, \hat F_n)) \\ = \alpha - P_F(n^{1/2}(\bar X_n - \theta(F))/\hat\sigma_n < z_\alpha + \frac 16\gamma(F)(z_\alpha^2 - 1)n^{-1/2}) + O(n^{-1})\]By similar reasoning, the two-sided bootstrap interval of nominal level $1-2\alpha$, given by
\[[\bar X_n - n^{-1/2}J_n^{-1}(1-\alpha, \hat F_n), \bar X_n - n^{-1/2}J_n^{-1}(\alpha, \hat F_n)]\]has coverage error $O(n^{-1})$
although these basic bootstrap intervals have the same order of coverage error as those based on the normal approximation, there is evidence that the bootstrap does provide some improvement (in terms of the size of the constants)
Bootstrap-t Confidence Intervals
consider bootstrap confidence intervals for $\theta(F)$ based on the studentized root
\[R_n^s(X^n, \theta(F)) = n^{1/2}(\bar X_n - \theta(F)) / \hat\sigma_n\,,\]the bootstrap versions of the expansions (15.46) and (15.47) are
\[K_n(t, \hat F_n) = \Phi(t) + \frac{1}{6} \gamma(\hat F_n)\phi(t) (2t^2+1) n^{-1/2} + O_P(n^{-1})\,, \tag{15.58}\]and
\[K_n^{-1}(1-\alpha, \hat F_n) = z_{1-\alpha} - \frac 16 \gamma(\hat F_n)(2z_{1-\alpha}^2+1)n^{-1/2} + O_P(n^{-1}) \tag{15.59}\]By comparing (15.46) and (15.58), it follows that
\[K_n(t, \hat F_n) - K_n(t, F) = O_P(n^{-1})\,,\]since $\gamma(\hat F_n) - \gamma(F) = O_P(n^{-1/2})$. Similarly,
\[K_n^{-1}(1-\alpha, \hat F_n) - K_n^{-1}(1-\alpha, F) = O_P(n^{-1})\]the bootstrap is more successful at estimating the distribution or quantiles of the studentized root than its nonstudentized version.
[TODO: omit details for the nominal level $1-\alpha$ upper confidence bound]
The one-sided coveraged error of the bootstrap-t interval is $O(n^{-1})$ and is of smaller order than that provided by the normal approximation or the bootstrap based on a nonstudentized root.
A heuristic reason why the bootstrap based on the root $R_n^s$ outperforms the bootstrap based on the root $R_n$ is as follows.
In the case of $R_n$, the bootstrap is estimating a distribution that has mean 0 and unknown variance $\sigma^2(F)$. The main contribution to the estimation error is the implicit estimation of $\sigma^2(F)$ by $\sigma^2(\hat F_n)$.
On the other hand, the root $R_n^s$ has a distribution that is nearly independent of $F$ since it is an asymptotic pivot.
Bootstrap Calibration
Let $I_n = I_n(1-\alpha)$ be any interval with nominal level $1-\alpha$. Its coverage is defined to be
\[C_n(1-\alpha, F) = P_F\{\theta(F)\in I_n(1-\alpha)\}\]We can estimate $C_n(1-\alpha, F)$ by its bootstrap counterpart $C_n(1-\alpha, \hat F_n)$. Then, determine $\hat\alpha_n$ to satisfy
\[C_n(1-\hat \alpha_n, \hat F_n) = 1 - \alpha\]To fix ideas, suppose $I_n(1-\alpha)$ is the one-sided normal theory interval $(-\infty, \bar X_n + n^{-1/2}\hat\sigma_nz_{1-\alpha})$.
\[C_n(1-\alpha, F) = P_F(n^{1/2}(\bar X_n - \theta(F))/\hat\sigma_n < z_\alpha) = 1-\alpha + \frac 16 \phi(z_\alpha)(2z_\alpha^2+1)n^{-1/2} + O(n^{-1})\]under smoothness and moment assumptions, the bootstrap estimated coverage satisifies
\[C_n(1-\alpha, \hat F_n) = 1-\alpha + \frac 16 \phi(z_\alpha) \gamma(\hat F_n)(2z_\alpha^2+1) n^{-1/2} + O_P(n^{-1})\,,\]and the value of $\hat\alpha_n$ is obtained by setting the estimated coverage equal to $1-\alpha$.
One can then show that
\[\hat\alpha_n -\alpha = -\frac 16\phi(z_\alpha) \gamma(F)(2z_\alpha^2+1)n^{-1/2} + O_P(n^{-1})\]Other Bootstrap methods
- iterate the calibration method
- Efron’s BC_a method
Comparison with Permutation Test
In the introduction of the section of Bootstrap method in Lehmann & Romano (2005), it said,
In the previous section, it was shown how permutation and randomization tests can be used in certain problems where the randomization hypothesis holds. Unfortunately, randomization tests only apply to a restricted class of problems. In this section, we discuss some generally used asymptotic approaches for constructing confidence regions or hypothesis tests.
So it sounds like the bootstrap procedure is more general than the permutation test, but I have not read the section about permutation test (TODO). Here I try to compare them in terms of test statistic.
To compare two group means with $\sigma_1=\sigma_2$, a natural test statistic is
\[t = \bar x_2 - \bar x_1\,.\](here the denominator $se$ is omitted) In the permutation test, we propose
\[t^*_p = \bar x_2^{*p} - \bar x_1^{*p} \,,\]while
\[t^*_b = (\bar x_2^{*b} - \bar x_2)- (\bar x_1^{*b} -\bar x_1)\]for bootstrap procedure, and the observations in the permutation test are sampled without replacement, while the observations in the bootstrap test are sampled with replacement. Detailed example can be found in bootstrap test for eigenfunctions and permutation test for gene.
More deep comparison with permutation test can be found in the one year ago’s post. I feel very guilty, I didn’t realize that I wrote such staff until I found there are two posts has the “bootstrap” tag.